
Aktuelles
Neuigkeiten und Veranstaltungen: Wir halten Sie auf dem Laufenden.
Aktuelle Meldungen
-
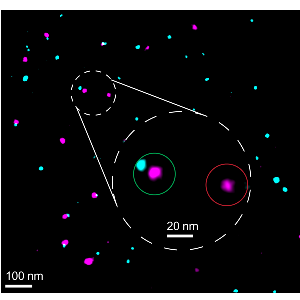 Testen, wie gut die Biomarker funktionieren
Testen, wie gut die Biomarker funktionierenLMU-Forschende haben eine Methode entwickelt, um bestimmen zu können, wie zuverlässig sich die Zielproteine in der superauflösenden Fluoreszenz-Mikroskopie markieren lassen.
-
Video: Professor Erwin Frey im Interview
Physiker Erwin Frey forscht über die Prinzipien des Lebens.
-
Wie entstehen Braune Zwerge?
Neue Beobachtungen geben Aufschluss darüber, ob die Geburt der Riesenplaneten ähnlich abläuft wie bei Sternen.
Aktuelle Veranstaltungen
-
 10 JunMünchner Physik Kolloquium
10 JunMünchner Physik KolloquiumVeranstaltungsreihe


